

 home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search
home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search


 home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search
home
overview
research
resources
outreach & training
outreach & training
visitors center
visitors center
search
search

Tom Goddard¹, Manfred Auer², A. James Hudspeth³, Wenyuan Shi4, and Mina J. Bissell²
¹
Resource for Biocomputing, Visualization, and Informatics
University of California, San Francisco
²
Life Sciences Division
Lawrence Berkeley National Laboratory
³
Laboratory of Sensory Neuroscience
Rockefeller University
4
Jonsson Comprehensive Cancer Center
University of California, Los Angeles
Studying Molecular Machines in the Native Cellular EnvironmentWith the completed sequencing of a variety of genomes and advances in proteomics, including high-throughput structural determination, the focus of modern biology is shifting towards the analysis of more complex systems such as cells and tissues. While biochemistry and structural biology traditionally focused on individual proteins, it has become clear over the last half a decade that the cellular proteome is highly organized, and that proteins are typically part of larger complexes and macromolecular machines, which are optimized to carry out particular cellular functions and which are highly regulated. As a consequence, the role of individual proteins needs to be understood in their native cellular context, if one aims to learn how their interactions underlie physiological function and how malfunction of single proteins gives rise to complex disease patterns. Therefore, visualization of protein complexes in cellular volumes in healthy and diseased tissue is a highly desirable yet daunting task.
Electron Microscope (EM) tomography is currently the only technique that allows the visualization of macromolecular machines in their native cellular habitat, as unlike all other structural approaches it does not rely on averaging of identical molecules. EM tomography is particularly well suited for proteins that are either too rare or fragile to withstand purification, or whose function depends on the complex interplay with other cellular components such as the cytoskeleton, the cell membrane or the extracellular matrix. EM tomographic imaging can provide 3-D protein envelopes at 3-5 nm resolution that - when the molecular composition is known - can be interpreted by docking candidate proteins into the density maps, yielding pseudo-atomic models of cellular volumes. EM tomography is also ideally suited as a discovery tool, as it allows an unbiased view into the molecular 3-D architecture of cells and tissues and can elucidate cellular differences between healthy and diseased tissue in unprecedented details.
Chimera: a Versatile Visualization Package for 3-D Cellular VolumesHowever, visualization of cellular volumes is far from trivial due to the size and complexity of the 3-D volume and the crowdedness of the molecular scenery. After experimenting with a variety of visualization packages such as Ono, PyMol, and Amira, we decided on Chimera due to its sophisticated tools for visualizing and analyzing 3-D density maps. During the course of the past year, we have been extensively interacting with Tom Goddard, resulting in a variety of additional visualization and analysis modules that have made Chimera all the more powerful.
Using some of these new features, we have been able to visualize 3-D volumes obtained by EM tomography, to extract features from these volumes, and to perform quantitative analysis such as exact 3-D length and volume measurements. We are very interested in continuing the collaboration with Tom Goddard and the RBVI to establish new tools that will allow interactive data exploration, using an area-of-interest multi-scale visualization, possibly virtual reality immersion visualization, on-the-fly-interactive region coding and 3-D segmentation, all of which would allow the rapid quantitative analysis of large tomographic data sets. In addition, I believe that Chimera is exceptionally well-positioned for correlative light and electron microscopy (multi-scale) imaging, which includes but is not limited to fluorescence light microscopy, EM serial section 3-D reconstruction, and EM tomography.
Sample Application: Molecular Understanding of our Senses of Hearing & BalanceOne in 1000 children is born deaf, and about 10% of the population is affected by severe hearing loss. Our senses of hearing and balance rely on the proper function of hair cells, epithelial cells of the inner ear that display a mechanosensitive organelle on their apical cell surface: the hair bundle. Hair bundles consists of several dozens of stereocillia that contain a highly cross-linked actin bundle core surrounded by plasma membrane and that are interconnected by at least three different kinds of extracellularly located filamentous linkers. The hair bundle is the location of mechano-electrical transduction, adaptation to sustained stimuli and also for amplification. In mammals, additional amplification is achieved by specializations of the lateral wall of outer hair cells.
Despite the clinical and biological importance of hair cells, we know very little about the exact molecular composition and 3-D architecture of the molecular machines that carry out these functions, in part due to the paucity and fragility of hair cells, which render classical biochemical and structural biology approaches futile. In addition, the mechanotransduction machinery consists of extracellularly located protein filaments, transmembrane complexes, including ion channels, as well as intracellularly-located motor proteins that are attached to the cytoskeleton. Therefore, this delicate hearing machinery requires its native cellular context to carry out its function and needs, therefore, to be studied in situ. Moreover, there is no reason to assume that each of these mechanotransduction, adaptation and amplification macromolecular complexes will be identical in every cell, as cellular processes are dynamic and protein-protein interactions are often transient. Yet, given the precision with which hair cells operate we expect an underlying conserved complex architecture that allows a robust response of these cells to mechanical signals.
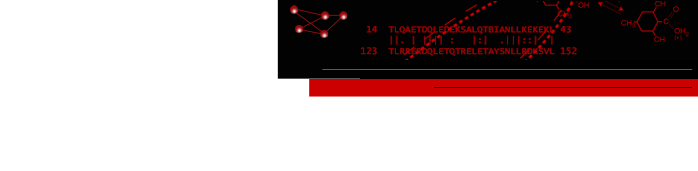
The mechanoelectrical transduction and adaptation machinery of hair cell stereocilia consists of extracellularly-located fine filaments that connect two adjacent stereocilia and whose stretching results in the direct opening of mechanoelectrical transduction channels without the involvement of chemical messengers. The direct nature of mechanoelectrical transduction implies that adaptation to sustained stimuli is achieved by adjustment of the tension in the tip links via a movement of a non-conventional myosin along the actin filament bundle. Employing EM tomography and quantitative analysis of extracellular protein linker densities, we characterized the properties of stereocilia ankle links, kinociliary links and tip links, the latter being intimately involved in mechanotransduction, and have concluded that the controversial implication of Cadherin 23 as the candidate for the long-sought after tip link is not consistent with our data of tip link length dimensions (Figure 1) [1,2]. This study indicates that EM tomography can discriminate among different candidate proteins that were proposed by various methods, as long as the candidates are sufficiently different in either shape or size. By docking actin bundle coordinates obtained by cryo-EM into tomographic reconstructions, we were able to validate the molecular preservation of our 3-D reconstruction. Furthermore, due to 3-D rendering and segmentation/differential coloring of map zones we were able to discover an auxiliary link that has not been reported so far. When viewing the 3-D volumes slice by slice, one easily misses this whole feature, emphasizing the need for interactive 3-D rendering of large and complex data sets. We will continue to use Chimera for a number of distinct molecular machines in the inner ear hair bundle, such as the complex adaptation machinery, as well as the stereocilia rootlet anchor, and also the Outer Hair Cell lateral wall amplifier. In addition, we have started to perform correlative LM & EM studies on Outer Hair Cells with the aim of localizing with high precision ion channels using biotinylated toxins followed by streptavidin-conjugated quantum-dots.
 Figure Caption: Hair cell.
Figure Caption: Hair cell.
Top left: Kinociliary links connecting stereocilia with kinocilium.
Middle left: Kinociliary links, gallery of individual kinociliary links.
Bottom left: Tip link with superimposed model.
Top right: Tip links and previously undescribed auxiliary links.
Bottom right: Atomic model of actin bundle obtained by cryo-EM docked into stereocilia actin core density envelopes, obtained by EMReferences:
- Gillespie PG, Dumont RA, Kachar B. (2005) "Have we found the tip link, transduction channel, and gating spring of the hair cell?" Curr Op Neurobiol 15:389-396.
- Auer M, Koster AJ, Ziese U, Bajaj C, Volkmann N, Wang DN & Hudspeth AJ. "3-D architecture of hair cell ankle, kinociliary and tip links as revealed by electron tomography." Submitted to J. Neuroscience.
- Kaplan HB. (2003) "Multicellular Development and gliding motility in Myxococcus xanthus." Curr Op Microbiol 6:572-577.
- Palsdottir H, Remis JP, Schaudinn C, Lux R, McDonald KL, Shi W, Costerton WJ & Auer M. "Visualization 3-D architecture of high-pressure frozen Myxococcus xanthus biofilms by EM tomography." In preparation.
- Bissell MJ, Radisky DC, Rizki A, Weaver VM, Peterson OW (2002) "The organizing principle: microenvironmental influences in the normal and malignant breast." Differentiation 70:537-546.
- Palsdottir H, Fournier M, Bissell MJ, Auer M. "Ultrastructural characterization of 3D cultured human mammary epithelial S1 and T4 cells." In preparation.
Laboratory Overview | Research | Outreach & Training | Available Resources | Visitors Center | Search