Tool: Profile Grid
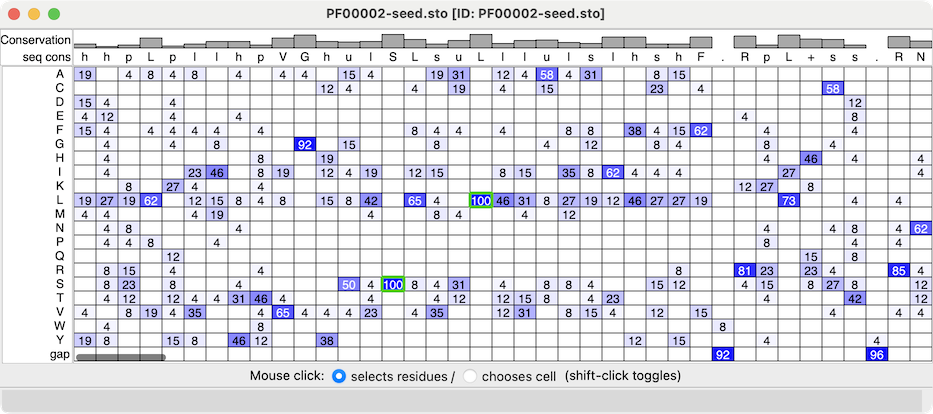
Profile Grid shows a condensed view of a multiple sequence alignment
in which the rows are the possible sequence characters (residue one-letter
codes and gap), the columns are positions in the alignment
(as in the standard view of a sequence alignment), and the values
in the cells are the prevalences of that type of residue at that position.
Profile Grid is a reimplementation of the viewer developed
by Alberto Roca, as described in:
ProfileGrids: a sequence alignment visualization paradigm that avoids the limitations of Sequence Logos.
Roca AI. BMC Proc. 2014 Aug 28;8(Suppl 2 Proceedings of the
3rd Annual Symposium on Biologica):S6.
By default,
the values in the ChimeraX Profile Grid are percentages
rounded to the nearest integer. A blank cell means that no sequences
at all have that residue type at that position, whereas a displayed value
of 0 means that some do, but <0.5%.
Much like the Sequence Viewer,
the Profile Grid tool interacts with any
associated structures, and it
can include headers above
the sequence data.
Typically one would use Profile Grid instead of the standard
Sequence Viewer
for alignments with very large numbers of sequences.
The sequence size threshold for determining the default viewer
can be adjusted in the
Sequences preferences,
although it can be overriden with the
open command
viewer option
(e.g., viewer grid
indicates using Profile Grid regardless of the alignment size).
Multiple Profile Grid windows can be shown at the same time,
and they can be manipulated like other panels in ChimeraX
(more...).
A Mouse click can interact with the grid cells in either of two ways,
as specified near the bottom of the window:
- selects residues
– selection of the corresponding
residue(s) in any associated
structures; if no associated structure has that type of residue at that
position, nothing happens.
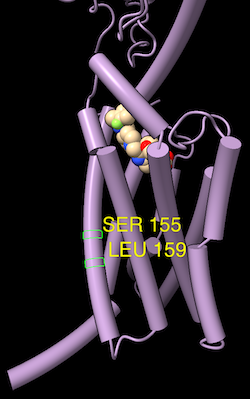
As in the Sequence Viewer,
selection of an associated structure residue
by any means will be shown as a green outline around the corresponding cell.
- chooses cell
– designating Profile Grid cells for subsequent actions,
shown with an orange diamond on each chosen cell.
Choosing a cell reports in the lower left corner of the window
the number of sequences matching that cell.
Either way, cell status can be toggled or the cell added to the existing set
of highlighted/chosen cells with Shift-click.
Moving the mouse cursor over a cell reports the column number in the
status area at lower right corner of the window, along with the number
of associated
structure chains (if any) and the residue type(s) in the
structure-associated sequences.
The residue type(s) in the structures could be different from those
in the sequences, but are usually the same
since association
uses the best match.
Context Menu
Settings
[back to top: Profile Grid]
Context Menu
The Profile Grid context menu
includes:
- Choose Cells for Sequence
▶ [individual sequence names]
– choose all cells corresponding to the
specified sequence
- Chosen Cells
– actions based on the chosen cells, if any:
- Change in Prevalence... bring up a dialog for showing
changes in residue-type prevalences for the chosen subset of sequences
relative to all sequences:
- Color other columns by prevalence change:
how to color columns without chosen cells
- Use [N] colors/thresholds (initial default 3)
– the fold-change values (coloring thresholds) can be edited directly,
and each color changed by clicking its color well and using the
system color editor
- Set colors from palette – change the colors collectively
by choosing a palette
from the pulldown menu
(initial default ^rdbu-3:
)
- Reverse – reverse the order of the colors
- But color cells with less than [percent] of sequences
originally [color]
(initial default on, color cells with original full-alignment prevalence
<0.5% in
)
- Color transitions:
- smooth (initial default) – between threshold values,
interpolate from one color to the next
- sharp – change color abruptly halfway between the
threshold values
- Color chosen cells [color] (initial default on and
)
- Color unchosen cells in chosen columns [color]
(initial default on and
)
- Revert dialog to the last-used settings
- Reset dialog to factory-default settings
- Remove prevalence coloring
- Apply the current settings
- List Sequence Names – list the names of the corresponding
sequences in a separate dialog for copying or saving to a text file
- New Alignment – open the corresponding sequences in a
new window in the specified viewer:
- Find Cell Pattern...
show dialog to find short stretches of conserved residue types generally
or that match a user-specified pattern:
- pattern [sequence-pattern]
– specify a sequence pattern with single-letter amino acid codes,
where case is not important. Multiple allowed types at a single
position can be given as a set of single-letter characters
within square brackets, or as a set of types to exclude within curly
brackets, or as a dot (period) to signify that all types are allowed.
Each allowed residue type must meet the
prevalence criterion.
For example, a search for RGD or RGE in three consecutive
columns of the alignment could be entered as rg[de], and for
CXXC could be entered as C..C, or to disallow cysteines
from the second and third positions, C{C}{C}C.
However, an important shortcoming is that the search does not allow for
variable spacing due to gaps, so that sequences with CXXC will not be
detected if the motif is spaced out to more than four columns
in the overall alignment.
- [N] consecutive non-gap cells
– find potential motifs in the alignment as
N consecutive residues of specific types that meet the
prevalence criterion
-
where each cell has [percent] occupancy or higher
– prevalence criterion
- Headers
– which headers are shown:
- Consensus
- Conservation
- RMSD – root-mean-square distance among residues
associated with a column,
based on Cα for peptides, C4' for nucleic acids
- Save ▶
[choice of available headers]
– save header information to file.
Headers are saved to a simple text format that lists the alignment positions
and values. See also:
sequence header
- Individual Sequence as Header
▶ [individual sequence names]
- Save Image... open browser dialog for saving the currently
visible part of the alignment to an image file, with settings:
- file name and location
- image file type (several choices, initial default PNG)
- View spacing (initial default 2)
– pixel spacing between the four sections
(header names, headers, residue types, and profile grid)
that will be composited together to create a single image
- DPI – image resolution in pixels per inch
- Scroll To Show New Selection If Needed (default on)
- Settings... show the Profile Grid
preferences to control cell contents
- Structure
- Associations... show dialog for controlling
sequence-structure association
-
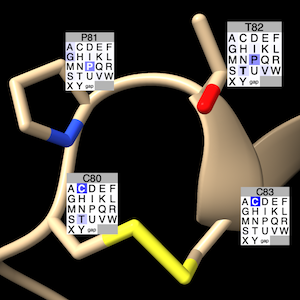
Label Residues... show dialog to label
associated structure residues
with grids color-coded by residue-type prevalence.
The structure residue's type is shown in the label title
(along with its number) and with bold font in the grid.
Settings:
- Chains – choose the structure chain of interest
- Also limit to selected residues, if any –
whether to limit labeling to selected
residues, or if none are selected, all residues in the chosen chain(s)
- Use [N] colors/thresholds – specify the
coloring palette. The default is from white to blue with increasing prevalence:
palette 0,white:1,blue
range 0.0,1.0
- Set colors from palette [palette-name]
– choose a built-in palette.
Only the ones with the same number of colors as the current number of
thresholds will be listed; for example, the rainbow palette will
not be listed unless there are exactly 5 thresholds.
If any colors are set interactively, the palette name will be
shown as custom.
- Reverse colors – reverse the order of the current
colors without changing the number of thresholds or their positions
- Background color
– color to use behind the label text and for filling in
empty cells in the last row of the label (default #B4B4B4
)
Clicking Apply (or OK, which also dismisses the dialog)
shows the labels, whereas Close simply dismisses the dialog.
Other buttons Revert the dialog to last-used settings, Reset
to factory defaults, and show Help in the
Help Viewer.
See also: sequence grid
- (Mac only)
Suppress Horizontal Scrollbar (default on)
– hide the horizontal scrollbar to prevent vertical misalignment
between rows and their labels; the window can still be scrolled horizontally,
e.g., with the trackpad
[back to top: Profile Grid]
Settings
Choosing Settings... from the
Profile Grid context menu
shows its settings in a separate window, with sections:
The settings window can be manipulated like other panels in the
ChimeraX interface (more...).
See also the Sequences
section of the main ChimeraX settings dialog.
Save saves the current settings as
preferences,
Reset replaces the current settings with
the initial “factory” defaults (values shown in bold below),
and Restore restores values that were saved previously.
The Buttons below... option indicates
whether these buttons should apply only
to the currently shown section (e.g., Appearance)
or to all of the Profile Grid preferences.
Although there can be multiple settings windows with different
values for multiple Profile Grid windows,
there can be only one set of saved preference values.
←
Appearance – grid cell contents
- Cell text
- none – no text, use color only to indicate prevalence
- count – raw counts
- percentage (initial default)
– counts as a percentage of the number of sequences
- Decimal places for percentages [N]
– for percentage values, the number of digits following the decimal
(initial default 0)
←
Headers – rows of information above the grid of residues
(more...)
UCSF Resource for Biocomputing, Visualization, and Informatics /
June 2026