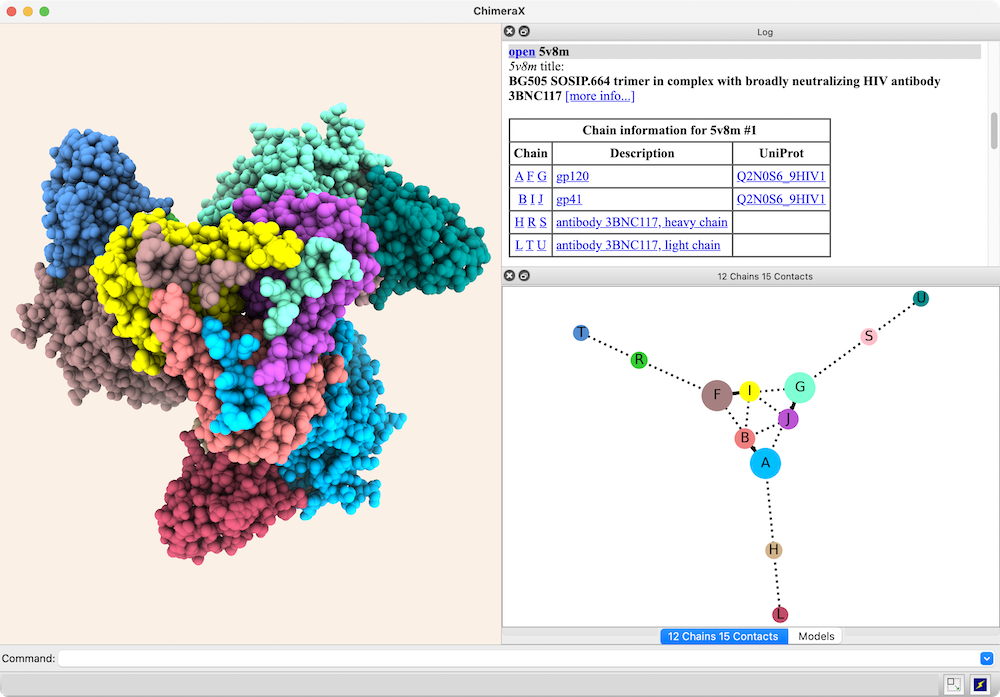
Chain-chain interfaces can be identified by buried surface areas
and displayed as a network diagram with the
interfaces
command or the
Molecule Display
icon
 .
In the diagram, nodes (circles) represent chains,
larger for greater surface areas,
and edges (lines) between nodes represent chain-chain interfaces
(default ≥ 300 Å2 buried area).
Dotted lines represent interfaces smaller than half the size of the
largest in the structure. Diagram context menus enable a variety of actions,
such as “exploding” the structure by moving chains apart,
hiding all but the chains in contact with a given chain,
and showing a more detailed plot of the residues forming a given interface.
.
In the diagram, nodes (circles) represent chains,
larger for greater surface areas,
and edges (lines) between nodes represent chain-chain interfaces
(default ≥ 300 Å2 buried area).
Dotted lines represent interfaces smaller than half the size of the
largest in the structure. Diagram context menus enable a variety of actions,
such as “exploding” the structure by moving chains apart,
hiding all but the chains in contact with a given chain,
and showing a more detailed plot of the residues forming a given interface.
The structure is an HIV envelope glycoprotein trimer
bound by three copies of a broadly neutralizing antibody
(PDB 5v8m), with chain information shown in the
Log.
Glycosylations (not displayed) were included in the surface area calculations.
For setup, see the command file
trimer-network.cxc.
More features...